Abstract
Within Europe, classical genetic markers, nuclear autosomal and Y-chromosome DNA polymorphisms display an east–west frequency gradient. This has been taken as evidence for the westward migration of Neolithic farmers from the Middle East. In contrast, most studies of mtDNA variation in Europe and the Middle East have not revealed clinal distributions. Here we report an analysis of dys44 haplotypes, consisting of 35 polymorphisms on an 8 kb segment of the dystrophin gene on Xp21, in a sample of 1203 Eurasian chromosomes. Our results do not show a significant genetic structure in Europe, though when Middle Eastern samples are included a very low but significant genetic structure, rooted in Middle Eastern heterogeneity, is observed. This structure was not correlated to either geography or language, indicating that neither of these factors are a barrier to gene flow within Europe and/or the Middle East. Spatial autocorrelation analysis did not show clinal variation from the Middle East to Europe, though an underlying and ancient east–west cline across the Eurasian continent was detected. Clines provide a strong signal of ancient major population migration(s), and we suggest that the observed cline likely resulted from an ancient, bifurcating migration out of Africa that influenced the colonizing of Europe, Asia and the Americas. Our study reveals that, in addition to settlements from the Near East, Europe has been influenced by other major population movements, such as expansion(s) from Asia, as well as by recent gene flow from within Europe and the Middle East.
Similar content being viewed by others
Introduction
The archaeological record suggests three main demographic expansions in Europe: the upper Paleolithic colonization, the Mesolithic re-expansion, following the last glacial maximum, and the Neolithic migration.1 The overall pattern of modern European genetic diversity should therefore reflect the joint effects of these three demographic expansions. The debate has arisen over the fraction of the modern European gene pool derived from each demographic event.2,3,4 Two major proposed models have resulted from the debate, which differ in their predictions of the origin of today's European gene pool: the demic-diffusion and cultural-diffusion models. The demic-diffusion model5 suggests that the spread of agricultural techniques to Europe was accompanied by the extensive immigration of Neolithic farmers from the Near East, who in consequence contributed a major fraction of the present-day European gene pool. In contrast, the cultural-diffusion model6 suggests that the transfer of agricultural technology occurred without significant population movement, such that most contemporary diversity within Europe would be rooted in the Paleolithic.
Geographical differentiation and clinal patterns in allele frequencies, extending from the Levant into Northern and Western Europe, were initially detected in studies of protein polymorphisms,7,8,9 and subsequently supported by nuclear autosomal and Y-chromosome DNA markers.10,11,12,13 These patterns were thought to have originated from a demic diffusion of Near Eastern farming communities into Europe in the early Neolithic period. On the contrary, most studies of mtDNA variation in Europe have shown less differentiation, and a lack of broad clinal variation.14,15,16,17,18 Initial analyses of mtDNA suggested that the majority of extant mtDNA lineages entered Europe during the Upper Palaeolithic.19 However, a detailed reanalysis of European mtDNA data did detect a gradient similar to that known from other markers,20 though it has been suggested that this gradient could have been established during the original Upper Palaeolithic colonization, and/or as a result of more recent (post-Neolithic) gene flow, rather than during the Neolithic expansion.20 MtDNA evidence has therefore not been consistent with the demic-diffusion model, but rather supported the cultural-diffusion model of the Neolithic agricultural revolution.
Studies of European genetic diversity have mostly been based on mitochondrial and Y-chromosome DNA. These marker systems offer the advantage of easily derived haplotypes, but only represent the genetic history of either females or males, respectively. Systematic studies of non-Y-chromosome nuclear DNA segments in European populations, reflective of the history of both males and females, are still lacking. To fill this gap, our group has characterized DNA polymorphisms in an 8 kb intronic segment, flanking exon 44 (dys44) of the human dystrophin gene on Xp21. Variation at this site has proved to be a very useful marker for human evolutionary studies,21 and led us to propose that two separately evolving lineages have contributed to the genetic diversity of modern humans.22 In the present study, we have analyzed the distribution of dys44 haplotypes in 19 European and Near Eastern populations within a broad Eurasian context, in order to identify the evolutionary and demographic events that shaped the contemporary European gene pool. We demonstrate that the distribution of X-chromosome variation in Europe and the Middle East lacks a strong geographic pattern, thus substantially differing from the clinal distribution observed with a variety of other markers. However, an underlying, ancient east–west gradient of X-chromosomal variation can be detected across the whole of Eurasia, and we suggest that this gradient has been disrupted by recent demographic events.
Materials and methods
Population samples
We analyzed a total of 1203 chromosomes from 19 Eurasian populations. Of these, 873 chromosomes from six European and seven Asian populations were previously characterized (Zietkiewicz et al, Am J Hum Genet 2003; 73).21,22,23 Recently, we analyzed a further 330 chromosomes of Mediterranean (39 Cretans), Iberian (32 Spanish) and Middle Eastern origin. The Middle Eastern samples were obtained from the National Laboratory for the Genetics of Israeli Populations at Tel-Aviv University, and included samples from Ashkenazi Jewish populations from Poland (24), Romania (seven) and Ukraine (seven), and Jewish populations from Bulgaria (18), Turkey (12), Morocco (29), Iran (21), Iraq (27) and Yemen (32), as well as three non-Jewish Arab populations: Druze (33), Palestinians (23) and Bedouin (26). The data on 70 Ashkenazi chromosomes analyzed earlier (Zietkiewicz et al, Am J Hum Genet 2003; 73) were also included. Given that population pairwise FST values indicated that the Jewish populations were not significantly different from one another based on dys44 haplotype frequencies (data not shown), and the relative lack of differentiation reported in the literature (eg Hammer et al24), all Jewish samples were treated as a single population. A list of the populations sampled, their geographic locations and linguistic affiliations are given in Table 1.
Typing dys44 polymorphisms
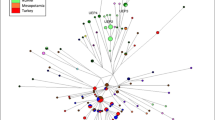
The 8 kb DNA dys44 segment consists of exon 44 (148 bp, between cDNA positions 6499 and 6646) and the surrounding intronic region (between position −2853 to −1 upstream and 1 to 5034 downstream) of the human dystrophin gene on Xp21 (accession number: U94396).21 In all, 35 intronic polymorphisms, including 32 single nucleotide substitutions (one three-allelic site), two three-nucleotide deletions and one eight-nucleotide duplication, were previously ascertained by SSCP/heteroduplex analysis of 7622 bp in 250 worldwide distributed chromosomes (Figure 1).23 New DNA samples were typed by allele-specific oligonucleotide (ASO) hybridization, as described by Zietkiewicz et al,23 to determine allelic states at the 35 segregating sites.
Eurasian diversity of the dys44 haplotype. The dys44 haplotype consists of 35 intronic polymorphisms, including 32 single-nucleotide substitutions (one three-allelic site), two three-nucleotide deletions, and one eight-nucleotide duplication (at position 3144). Positions of each site are shown in numbers of bases up or downstream from exon 44 of the human dystrophin gene. Ancestral alleles are given for each site at the top of the diagram and the derived states for each of the 60 observed haplotypes are listed; the blank spaces indicate the presence of the ancestral state.
Haplotype derivation and statistical analyses
Unambiguous dys44 haplotypes were inferred directly from genotypes in hemizygous males, homozygous females and females heterozygous at only one position. Haplotypes for females heterozygous at multiple positions were reconstructed from the genotype data as in Labuda et al,22 and confirmed by the program PHASE.25
Gene diversity, nucleotide diversity, Ewens's θ, genetic distances (as a pairwise FST matrix) and the analysis of molecular variance (AMOVA)26 were obtained using ARLEQUIN Version 2.0.27 FST matrices were computed with haplotype frequencies across populations (results were unaffected when FST matrices were calculated, also taking into account interhaplotype distances). Population comparisons were performed through a multidimensional scaling (MDS) analysis of the pairwise FST distances with Statistica 6.0 (StatSoft, Inc.).
To quantitatively examine the geographic differentiation of dys44 haplotypes, spatial autocorrelation analyses were carried out using the program AIDA, specifically designed for DNA analysis.28 Analyses were undertaken with DNA diversity data using all haplotypes, or only the six most frequent haplotypes. AIDA measures the overall genetic similarity between population samples at arbitrary geographic distance classes and assesses significance by a series of permutation tests. In this study, equal-interval distance classes were used, and the first distance class (0) included comparisons between haplotypes belonging to the same population. Similarity between haplotypes, or positive autocorrelation, is shown by positive II values, while haplotype dissimilarity, termed negative autocorrelation, results in negative values of II. A set of spatial autocorrelation coefficients (designated as II) evaluated at various distance classes (termed the correlogram) can be associated with one or more demographic processes.28,29
The relative importance of language affiliation and geography in the shaping of genetic diversity was assessed by calculation of the coefficients between genetic, linguistic and geographic distance matrices, whose levels of significance were evaluated by Mantel test30 implemented in the ARLEQUIN package. The geographic distances were entered as a matrix of the great-circle distances between pairs of populations (following the method of Rousset,31 we also repeated the Mantel test with the logarithm of the great-circle distances; results were consistent across both methods), assessed on the basis of population geographic coordinates (Table 1) obtained from the World Atlas online database (http://www.worldatlas.com). The matrix of linguistic distances was built as described previously by Excoffier et al32 and Poloni et al33 Briefly, a dissimilarity index of 8 was arbitrarily assigned to any pair of populations belonging to different language families. Pairs of populations of the same language family (for example Indo-European), but of different subfamilies (for example Germanic and Slavic), were assigned a dissimilarity index of 3. This distance was decreased by 1 for each shared level of classification (as defined by the Ethnologue online language database), up to three shared levels, where the distance was set to 0. A neighbor-joining tree showing the arbitrary distances assigned between groups according to their linguistic relationships is shown in Figure 5c. The linguistic classification of world languages used in this process was adopted from Ruhlen34 and the Ethnologue online language database (http://www.ethnologue.com/family_index.asp).
Population pairwise comparisons through MDS analysis based on the population pairwise FST distance matrix for all sampled populations (a), and for European and Middle Eastern populations (b). Different symbols code for linguistic affiliations. To allow comparison between the genetic and linguistic structure, a neighbor-joining tree showing the arbitrary distances assigned between groups according to their linguistic classification, for purposes of the Mantel Test, is also shown (c).
Results
Geographical distribution of dys44 haplotypes in Eurasia
In total, 60 distinct dys44 haplotypes were seen in 1203 X-chromosomes from 19 Eurasian populations (Figure 1). The six most frequent haplotypes (B001, B002, B003, B005, B006 and B008) represented 88% of all analyzed chromosomes. The remaining 54 haplotypes tended to be either rare or absent from certain populations. The frequency distributions of these six most frequent dys44 haplotypes are shown in Figure 2. Each of the haplotypes displayed a relatively even distribution from the Middle East to Europe, emphasizing the genetic similarity between these regions. In contrast, the comparison of haplotype frequencies in Europe and the Middle East on the one hand, and Central and Eastern Asia on the other, revealed large differences in the frequency distributions of haplotypes B002 and B003.
The most frequent haplotype B001 occurred at a very high frequency in all Eurasian populations (from 0.26 to 0.60), except for the Basques, where its frequency was only 0.15. In Asia, an eastward increase of B001 was observed, from 0.26 in a Mongolian group (Oirat1), up to its highest frequency of 0.60 in the Japanese. The distribution of B002, the second most frequent haplotype, revealed pronounced geographic differences. It was rare in Europe (0.06) and the Middle East (0.06), but relatively frequent in Asia (0.26). In contrast to B002, the third most frequent haplotype B003 shows an opposite distribution, being relatively frequent in Europe (0.23) and the Middle East (0.19), but rare in Asiatic populations (0.06).
The other three most common haplotypes, B005, B006 and B008, were observed at a relatively low frequency. B006 was seen in all populations from Europe and the Middle East, except for the Bedouin, and was very rare in Asia. B005 was present in all Asians and in most populations from Europe and the Middle East, while B008, the least frequent among the six most common haplotypes, ranged from an average of 0.01 in Asia to 0.07 in the Middle East.
Gene diversity of dys44 haplotypes in Eurasia
All Eurasian populations exhibited high levels of gene diversity, which varied from 0.890 in the Basque to 0.569 in the Japanese (Table 1). An eastward decrease of gene diversity was observed: the average gene diversity ranged from 0.812 in Europe and 0.766 in the Middle East to 0.721 in Asia; within Asia, it decreased from 0.826 in Siberians to 0.569 in the Japanese. The average nucleotide diversities were 7.1 × 10−4 in Europe, 6.6 × 10−4 in the Middle East and 8.5 × 10−4 in Asia; the highest value of nucleotide diversity (9.9 × 10−4) was observed in a Mongolian group (Oirat1), and the lowest (3.4 × 10−4) in the Bedouin.
Ewens's method,35 based on the infinite-allele model, was used to estimate the population parameter θ. The Bedouin and Japanese presented notably lower θ values (Table 1), which could be due to a long-term smaller effective population size or founder effects. To ensure that the Bedouin did not affect the spatial autocorrelation analyses described below, they were removed and the analyses repeated; their removal did not affect the AIDA profile (data not shown).
Analysis of geographic structure by spatial autocorrelation
In order to quantitatively examine the geographic differentiation of dys44 haplotypes throughout Eurasia, we performed spatial autocorrelation analyses28 both by comparison of DNA sequence similarity using the entire haplotype dataset, and using frequencies of each of the six most common dys44 haplotypes B001, B002, B003, B005, B006 and B008, respectively (Figure 3). The pattern based on the entire haplotype dataset is strongly clinal, in which the autocorrelation coefficients (II) decrease from being significantly positive to significantly negative with increasing geographic distance. The II value is positive and highly significant at distance 0, which indicates that genetic similarity is higher within, than between population samples. Coefficients are positive and significant for all distances <4000 km, and are negative and significant for all distances >4000 km, essentially differentiating between Europeans and Asians. However, the coefficients show an upward trend for the 2000–3000 km distance class. Fluctuations of this kind are referred to as ‘long-distance differentiation’28 and suggest a recent disturbance of the cline, though they could also simply be the result of the discontinuous distribution of population samples. The coefficients also show an upward trend for the two longest distance classes (>9000 km), consisting of comparisons between the Japanese and most European and Middle Eastern samples, which suggests that the basic clinal pattern is restricted to the continental mainland.
Frequencies of haplotypes B002 and B003, which together account for 30% of all sampled Eurasian chromosomes, show a similar, continental scale clinal pattern from Asia to Europe (Figure 3). While haplotype B005 shows a clinal distribution in the short-distance classes (<6000 km), the coefficients become positive or zero for distances >7000 km (a ‘depression’), possibly indicating a regionally localized cline caused by phenomena affecting only part of the continent (though, as noted above, fluctuations in a correlogram's profile could simply be due to sample distribution). No clinal variation was observed for haplotypes B001, B006 and B008.
To test for the geographic differentiation of dys44 within Europe and the Middle East, a spatial autocorrelation analysis was performed with only the European and Middle Eastern samples (Figure 4). The resulting correlograms show a lack of clinal variation and all the coefficients are insignificant across all distance classes. Each of the six most frequent haplotypes within Europe and the Middle East also show a spatially random distribution of variation. To allow more population comparisons at each distance class, and hence a greater chance of observing any clinal variation present, we repeated the analysis using five (instead of ten) distance classes across Europe; the results remained insignificant and no cline was observed (data not shown).
Test of population structure by AMOVA
With all Eurasian populations treated as a single group, an AMOVA estimated that only 5.5% of the total genetic variance was due to differences among populations (i.e. FST=0.055; Table 2); in three groups (Europe, the Middle East and Asia), 5.88% of the genetic variance was due to differences between groups, and when in two groups (Europe/Middle East and Asia) differences between groups increased to 7.94% of the variance. This indicates that the structure of Eurasian X-chromosome diversity was mainly caused by differences between Europe/Middle East and Asia.
Considering only the European and Middle Eastern samples as one group, the FST value was low but significant (Table 2); however, when the four Middle Eastern populations were excluded, the genetic variance among the European populations alone was insignificant. These results highlight the lack of variation within Europe. When the European and Middle Eastern samples were divided into two groups, Europe and the Middle East, the difference between groups was insignificant, but that among populations within groups was significant. This, together with the previous AMOVA results, indicates that European and Middle Eastern populations are not genetically differentiated, and that any visible genetic structure across Europe/Middle East is due to variation among the four Middle Eastern populations.
Population comparisons through MDS analysis
Relationships among population samples were described by MDS analysis based on the pairwise FST distance matrix. When all populations were analyzed (Figure 5a), the major division was found between Asia and Europe/Middle East; the Asian samples clustered in the upper right corner of the plot, and were quite distinct from the Europeans and Middle Easterners. In the Asian cluster, three Mongolian groups, the Turkic, Khalkha and Oirat2, displayed close affinity, and were well separated from another Mongolian group, Oirat1. The Siberians were relatively close to the remaining Asians, but also showed affinities with Europe and the Middle East.
European and Middle Eastern populations were not well separated on the plot. Rather, the Europeans clustered tightly in the center, surrounded by the Middle Eastern populations, indicating that the Middle East is more heterogeneous than Europe. The Bedouin appeared distinct from other Middle Easterners and Europeans, indicating that a bottleneck and/or founder effect, followed by genetic drift, had a strong influence on the dys44 haplotype distribution of this small population. A second small and distinct population is the Basques, who speak a non-Indo European language, seemingly unrelated to any other languages. This linguistic isolation seemed to be reflected in the MDS analysis, in which the Basques were somewhat distinct from other Europeans.
European and Middle Eastern populations were also not separated in the MDS analysis of just Europe/Middle East (Figure 5b). As in the MDS analysis of the whole of Eurasia, the Middle Eastern samples surrounded a relatively less diverse cluster of European populations. It is apparent that there is some degree of genetic continuity between European and Middle Eastern populations, and little evidence of clustering by either geography or language.
Correlation tests between genetic, geographic, and linguistic distances
Mantel tests30 were used to assess the relative importance of different factors in the shaping of genetic diversity (Table 3). We calculated correlation coefficients between pairs of factors (genetics, language and geography), and partial correlation coefficients between genetics and language, and between genetics and geography, with the third factor kept constant. For the Eurasian samples, we found that the correlations between genetics and language, and genetics and geography, were strong and significant (in both cases P<0.001). The partial correlation of genetics and geography was found to be less strong but still significant (P<0.01), while the partial correlation of genetics and language was low and insignificant (P>0.05). This analysis, therefore, confirms the primacy of geography, rather than language, in shaping the patterns of dys44 diversity within the Eurasian continent. All correlation and partial correlation tests performed for just Europe and the Middle East were insignificant (P>0.05).
Discussion
Although European populations have been intensively studied with a number of DNA markers, there is still no consensus on whether there is a clear population structure within the continent. The present study analyzed the patterns of X-chromosomal dys44 diversity of populations from Europe. In addition, the neighboring regions of the Middle East and Asia, which have played a role in the shaping of the European gene pool, were sampled. Our aim was to identify patterns of X-chromosome diversity, and infer from them the possible demographic and evolutionary events that, together, have shaped the European gene pool.
When taken together, the European and Middle Eastern samples showed a low but still significant X-chromosome genetic structure. However, this genetic structure did not reflect either a geographical or linguistic pattern within Europe. On the other hand, an underlying east–west clinal pattern of variation between Europe and Asia was detected. Here we discuss (i) what evolutionary factors could have contributed to the formation of these patterns, and (ii) what processes could have caused the discrepancies between the patterns identified at the X-chromosome region used in this study, and other molecular markers previously analyzed in European and Middle Eastern populations.
X-chromosome dys44 diversity in Europe: lack of geographic pattern
The AIDA analysis indicated that there is a remarkable lack of geographic structure in Europe (Figure 4); a finding confirmed by MDS analysis and AMOVA, which showed that Europeans could not be clearly distinguished from Middle Easterners (Figure 4). The lack of a clear overall pattern of the X-chromosome dys44 haplotype distribution in this study suggests that there was either never any cline of dys44 haplotypes (as discussed below) or that recent (post-Neolithic), substantial gene flow within Europe8 may have erased any previously existing signatures of population migration. Moreover, the MDS analysis and Mantel tests also suggest that neither language nor geography provides a strong barrier against gene flow in Europe.
The lack of observed geographic structure in X-chromosome diversity in Europe resembles that of European mtDNA diversity18 (though see Richards et al20). However, many other previously studied genetic markers, including protein, Y-chromosome and autosomal DNA markers, have all described east–west gradients suggestive of immigration from the Near East.7,9,10,11,12,13 Moreover, analyses of classical and Y-chromosome marker systems in Europe have shown a significant correlation between genetic and geographic distance, and somewhat less of a correlation between genetics and language affiliation.12,36 Therefore, the patterns of X-chromosome diversity in Europe presented here appear to differ from those observed by a number of previous studies using other marker systems.
Such discrepancies may be due to a number of factors. Firstly, selection is often cited as a reason for differences in allele distributions across loci.37 However, the dys44 segment lies in an intronic region in the middle of the large (2.4 Mb) dystrophin gene, and has a level of genetic diversity typical of neutral variation, and satisfies neutral expectations when tested with Tajima's38 D parameter.21 Furthermore, dys44 is found in a region of the X-chromosome that experiences a high recombination rate,39 and is thus very unlikely to be affected by selection on the neighboring loci. Secondly, because of the X-chromosome's mode of inheritance, the geographic structure of its genetic diversity has been relatively more affected by female, rather than male, migration. A recent (post-Neolithic) high rate of female mobility, suggested by Seielstad et al,40 may have erased any ancient clinal distribution of X-chromosome markers, while leaving the clinal patterns of autosomes and the Y-chromosome markers relatively unaffected. Given that the mitochondrial genome is only maternally transmitted, the high rate of female gene flow in Europe would have also strongly affected the pattern of mtDNA distribution. Indeed, most studies of mtDNA in Europe have revealed a lack of clinal variation.14,15,17,18 It is therefore likely that the higher rate of female gene flow in Europe is a major cause of the different patterns of X-chromosome markers when compared to autosomal and Y-chromosome DNA. Thirdly, and as suggested by Sokal et al,41 the absence of clines at certain loci can be fully consistent with the demic-diffusion model, as only loci at which haplotype frequencies were markedly different for pre-Neolithic European hunter gatherers and Neolithic farmers are expected to show clinal variation across Europe.5 The common variant dys44 haplotypes are old and were present in the ancestral, pre-out-of-Africa migration, and hence it is possible that allele frequencies did not vary between the incumbent and immigrating populations. Finally, the limited number of population samples available may pose a limiting factor to the detection of genetic structure in Europe. The low number of Middle Eastern and southern European samples, and the fact that one of these is from Crete, an island population with the inevitable consequences of drift due to its small size and relative isolation, suggests that even if a cline was to exist it would be hard to identify it with the current data set. Hence, it is possible that a westward gradient from the Near East to Europe could be found by analyzing a larger number of samples.
Traces of population expansion in Eurasia
MDS analysis and AMOVA, based on haplotype frequencies, showed that X-chromosome variation in the whole Eurasian continent is, in contrast to Europe alone, relatively well structured (FST=0.055, P<0.001). In particular, there was a significant division found between Europe/Middle East and Asia, and spatial autocorrelation analyses detected an underlying east–west gradient of X-chromosomal variation from Asia to Europe/Middle East. This observed cline remained significant even when European or Middle Eastern samples were removed from the data set (data not shown), showing that it is not specifically between Asia and Europe or Asia and the Middle East, but rather across the whole Eurasian continent. The ‘long-distance differentiation’ observed at the 2000–3000 km distance class suggests that the east–west cline has been disrupted by recent evolutionary and demographic events, such as successive gene flow, drift and/or adaptation to local environmental factors.28 Alternatively, the fluctuation in the correlogram profile could be due to the discontinuous distribution of population samples, though Y-chromosome microsatellites and binary markers have previously suggested a Central Asian genetic landscape reshaped by recent population events.42
Clines are an expected consequence of major population movements in which population expansion into new territories is accompanied by repeated founder effects and subsequent population growth.43,44 To be able to cause such a significant cline, it is necessary that the population expansion happened when population numbers were relatively small, and hence an expansion would be able to have a major effect on allele frequencies across a whole subcontinent. This reasoning, in accordance with the fact that the dys44 marker system has an increased time depth to that of mtDNA and Y-chromosome data, leads us to the conclusion that the demographic events that we witness through the cline were ancient ones (pre-Neolithic). However, while we suggest that we are witnessing Palaeolithic events, the ubiquity of the haplotypes involved means that we are unable to confidently date the time of the migration. Nevertheless, since the two haplotypes that contributed most strongly to the cline, B002 and B003, are found worldwide, including Sub-Saharan Africa, we can assume that they existed in the ancestral populations before the out-of-Africa expansion(s), and hence were likely to have been present in the initial colonization events. Therefore, the observed clinal distributions could suggest an ancient population expansion, probably from Asia to Europe, accompanied by repeated founder effects within the Eurasian continent. Alternatively, the distribution could be due to differences established during the initial colonization of Eurasia by anatomically modern humans during the Paleolithic, though the two are not necessarily mutually exclusive.
If an ancient population expansion within Eurasia was the cause of the observed cline, it can be argued, given the relative times of colonization of the two regions, that it was due to movement in a westerly direction from Asia to Europe. The expansion of anatomically modern humans out of Africa likely resulted in the colonization of Asia from the Near East and/or through the horn of Africa around 60 000 years ago, and only later of Europe through Anatolia around 35 000 years ago.8,45 It is therefore possible that the early Paleolithic populations in Asia moved into Europe through Central Asia, causing the observed cline, and thus modern Europeans might have received genetic contributions from two distinct Paleolithic populations: the Middle East and Asia (as shown by the gray arrows in Figure 6).
Schematic showing hypothetical routes of ancient migrations that could have caused the observed east–west Eurasian cline. Gray arrows: show an initial southern route leading to the colonization of southern Asian and Oceania (a), followed by an east–west migration towards the colonization of Europe, contemporary to the parallel colonization of Europe from Africa via the Middle East (b). Black arrows: show the out-of-Africa route leading to an ancestral population (1), from which bifurcating migrations lead to gene flow towards the Americas and Europe (2). Note that the migration patterns are not mutually exclusive.
Alternatively, the clinal variation could be due to a bifurcating colonizing event that initially occurred though the northern, out-of-Africa route, as described in the literature.45,46 It has previously been postulated from both mtDNA and Y-chromosome data that Europe and the Americas at least partly share a common ancestral gene pool. 47,48,49 The notion of a relationship between the early new world and Palaeolithic European populations has further been supported by the craniofacial data of Brace et al.50 A proposed ancestral population to both continents could have spread through the northern route and inhabited the Lake Baikal47,49 or Altaian48,51 regions of northern Central Asia, and from there expanded to both the Americas to the east and Europe to the west. MtDNA haplogroup ‘Brown's’ X,52 Y-chromosome haplogroup 1C47 and Y-chromosome haplotype 10 from Santos et al48 have all offered support to this hypothesis, and together bring to light the genetic similarities between Europe and the Americas. In addition, X-chromosome data (unpublished) from our laboratory also suggest a relationship between Europe and the Americas. Hence, it can be argued that the X-chromosome clinal variation observed across Eurasia is due, at least in part, to the migration of these ancient population groups (as shown by the black arrows in Figure 6). This ancient bifurcating event, resulting in populations with a common ancestry moving east and west across the northern fringes of Eurasia, and admixing with Asian populations there as a result of an earlier, southern out-of-Africa route, could conceivably have created the ancient clinal pattern that we can still observe today with deep time-depth markers such as dys44.
In conclusion, this study suggests that the demographic history of Europe, in addition to Neolithic expansions from the Near East, has been influenced by other major population movements, such as population expansions from Asia, and further reshaped by intracontinental gene flow. A large-scale survey of Eurasian X-chromosomal diversity would greatly assist in the identification of a number of migrations that have shaped the contemporary European gene pool, and provide more clues to understand the ancient routes and migrations of modern humans within Eurasia.
References
Barbujani G, Bertorelle G : Genetics and the population history of Europe. Proc Natl Acad Sci USA 2001; 98: 22–25.
Richards M, Corte-Real H, Forster P et al: Paleolithic and neolithic lineages in the European mitochondrial gene pool. Am J Hum Genet 1996; 59: 185–203.
Cavalli-Sforza LL, Minch E : Paleolithic and Neolithic lineages in the European mitochondrial gene pool. Am J Hum Genet 1997; 61: 247–254.
Barbujani G, Bertorelle G, Chikhi L : Evidence for Paleolithic and Neolithic gene flow in Europe. Am J Hum Genet 1998; 62: 488–492.
Ammerman AJ, Cavalli-Sforza LL : The Neolithic Transition and the Genetics of Populations in Europe. Princeton: Princeton University Press, 1984.
Dennell R : European Economic Prehistory: A New Approach. London: Academic, 1983.
Menozzi P, Piazza A, Cavalli-Sforza L : Synthetic maps of human gene frequencies in Europeans. Science 1978; 201: 786–792.
Cavalli-Sforza LL, Menozzi P, Piazza A : The History and Geography of Human Genes. Princeton, NJ: Princeton University Press, 1994.
Piazza A, Rendine S, Minch E, Menozzi P, Mountain J, Cavalli-Sforza LL : Genetics and the origin of European languages. Proc Natl Acad Sci USA 1995; 92: 5836–5840.
Chikhi L, Destro-Bisol G, Bertorelle G, Pascali V, Barbujani G : Clines of nuclear DNA markers suggest a largely neolithic ancestry of the European gene pool. Proc Natl Acad Sci USA 1998; 95: 9053–9058.
Chikhi L, Destro-Bisol G, Pascali V, Baravelli V, Dobosz M, Barbujani G : Clinal variation in the nuclear DNA of Europeans. Hum Biol 1998; 70: 643–657.
Rosser ZH, Zerjal T, Hurles ME et al: Y-chromosomal diversity in Europe is clinal and influenced primarily by geography, rather than by language. Am J Hum Genet 2000; 67: 1526–1543.
Semino O, Passarino G, Oefner PJ et al: The genetic legacy of paleolithic homo sapiens sapiens in extant europeans: A Y chromosome perspective (in process citation). Science 2000; 290: 1155–1159.
Torroni A, Lott MT, Cabell MF, Chen YS, Lavergne L, Wallace DC : mtDNA and the origin of Caucasians: identification of ancient Caucasian-specific haplogroups, one of which is prone to a recurrent somatic duplication in the D-loop region. Am J Hum Genet 1994; 55: 760–776.
Sajantila A, Lahermo P, Anttinen T et al: Genes and languages in Europe: an analysis of mitochondrial lineages. Genome Res 1995; 5: 42–52.
Comas D, Calafell F, Mateu E, Perez-Lezaun A, Bosch E, Bertranpetit J : Mitochondrial DNA variation and the origin of the Europeans. Hum Genet 1997; 99: 443–449.
Richards MB, Macaulay VA, Bandelt HJ, Sykes BC : Phylogeography of mitochondrial DNA in western Europe. Ann Hum Genet 1998; 62: 241–260.
Simoni L, Calafell F, Pettener D, Bertranpetit J, Barbujani G : Geographic patterns of mtDNA diversity in Europe. Am J Hum Genet 2000; 66: 262–278.
Richards M, Macaulay V, Hickey E et al: Tracing European founder lineages in the near eastern mtDNA pool. Am J Hum Genet 2000; 67: 1251–1276.
Richards M, Macaulay V, Torroni A, Bandelt HJ : In search of geographical patterns in European mitochondrial DNA. Am J Hum Genet 2002; 71: 1168–1174.
Zietkiewicz E, Yotova V, Jarnik M et al: Genetic structure of the ancestral population of modern humans. J Mol Evol 1998; 47: 146–155.
Labuda D, Zietkiewicz E, Yotova V : Archaic lineages in the history of modern humans. Genetics 2000; 156: 799–808.
Zietkiewicz E, Yotova V, Jarnik M et al: Nuclear DNA diversity in worldwide distributed human populations. Gene 1997; 205: 161–171.
Hammer MF, Redd AJ, Wood ET et al: Jewish and Middle Eastern non-Jewish populations share a common pool of Y-chromosome biallelic haplotypes. Proc Natl Acad Sci USA 2000; 97: 6769–6774.
Stephens M, Smith NJ, Donnelly P : A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 2001; 68: 978–989.
Excoffier L, Smouse PE, Quattro JM : Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 1992; 131: 479–491.
Schneider S, Roessli D, Excoffier L : Arlequin: A Software for Population Genetics Data Analysis. Switzerland: Genetics and Biometry Laboratory, Department of Anthropology, University of Geneva, 2000.
Bertorelle G, Barbujani G : Analysis of DNA diversity by spatial autocorrelation. Genetics 1995; 140: 811–819.
Sokal RR, Oden NL : Spatial autocorrelation in biology. Biol J Linn Soc 1978; 10: 199–249.
Mantel N : The detection of disease clustering and a generalized regression approach. Cancer Res 1967; 27: 209–220.
Rousset F : Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 1997; 145: 1219–1228.
Excoffier L, Harding RM, Sokal RR, Pellegrini B, Sanchez-Mazas A : Spatial differentiation of RH and GM haplotype frequencies in Sub-Saharan Africa and its relation to linguistic affinities. Hum Biol 1991; 63: 273–307.
Poloni ES, Semino O, Passarino G et al: Human genetic affinities for Y-chromosome P49a,f/TaqI haplotypes show strong correspondence with linguistics. Am J Hum Genet 1997; 61: 1015–1035.
Ruhlen M : A Guide to the World's Languages. London: Edward Arnold, 1987.
Ewens WJ : The sampling theory of selectively neutral alleles. Theor Popul Biol 1972; 3: 87–112.
Sokal RR : Genetic, geographic, and linguistic distances in Europe. Proc Natl Acad Sci USA 1988; 85: 1722–1726.
Cavalli-Sforza LL : Population structure and human evolution. Proc R Soc Lond B Biol Sci 1966; 164: 362–379.
Tajima F : Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 1989; 123: 585–595.
Oudet C, Hanauer A, Clemens P, Caskey T, Mandel JL : Two hot spots of recombination in the DMD gene correlate with the deletion prone regions. Hum Mol Genet 1992; 1: 599–603.
Seielstad MT, Minch E, Cavalli-Sforza LL : Genetic evidence for a higher female migration rate in humans. Nat Genet 1998; 20: 278–280.
Sokal RR, Harding RM, Oden NL : Spatial patterns of human gene frequencies in Europe. Am J Phys Anthropol 1989; 80: 267–294.
Zerjal T, Wells RS, Yuldasheva N, Ruzibakiev R, Tyler-Smith C : A genetic landscape reshaped by recent events: Y-chromosomal insights into central Asia. Am J Hum Genet 2002; 71: 466–482.
Cavalli-Sforza LL, Menozzi P, Piazza A : Demic expansions and human evolution. Science 1993; 259: 639–646.
Barbujani G, Pilastro A, De Domenico S, Renfrew C : Genetic variation in North Africa and Eurasia: neolithic demic diffusion vs Paleolithic colonisation. Am J Phys Anthropol 1994; 95: 137–154.
Lahr MM, Foley RA : Multiple dispersals and modern human origins. Evol Anthropol 1994; 3: 48–60.
Lahr MM, Foley RA : Towards a theory of modern human origins: geography, demography, and diversity in recent human evolution. Am J Phys Anthropol 1998; 27 (Suppl): 137–176.
Karafet TM, Zegura SL, Posukh O et al: Ancestral Asian source(s) of new world Y-chromosome founder haplotypes. Am J Hum Genet 1999; 64: 817–831.
Santos FR, Pandya A, Tyler-Smith C et al: The central Siberian origin for native American Y chromosomes. Am J Hum Genet 1999; 64: 619–628.
Wells RS, Yuldasheva N, Ruzibakiev R et al: The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc Natl Acad Sci USA 2001; 98: 10244–10249.
Brace CL, Nelson AR, Seguchi N et al: Old World sources of the first New World human inhabitants: a comparative craniofacial view. Proc Natl Acad Sci USA 2001; 98: 10017–10022.
Malhi RS, Smith DG : Brief communication: Haplogroup X confirmed in prehistoric North America. Am J Phys Anthropol 2002; 119: 84–86.
Brown MD, Hosseini SH, Torroni A et al: mtDNA haplogroup X: an ancient link between Europe/Western Asia and North America? Am J Hum Genet 1998; 63: 1852–1861.
Acknowledgements
We are grateful to all DNA donors who made this study possible. Tina Wambach provided many valuable comments on a draft manuscript. The manuscript was significantly improved, thanks to comments from three anonymous reviewers. This work was supported by a grant (# MOP-12782) from the Canadian Institutes of Health Research to DL. FXX was a recipient of a fellowship from the Research Center of Sainte-Justine Hospital.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xiao, FX., Yotova, V., Zietkiewicz, E. et al. Human X-chromosomal lineages in Europe reveal Middle Eastern and Asiatic contacts. Eur J Hum Genet 12, 301–311 (2004). https://doi.org/10.1038/sj.ejhg.5201097
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201097
Keywords
This article is cited by
-
Genetic polymorphisms of 10 X-STR among four ethnic populations in northwest of China
Molecular Biology Reports (2012)
-
Tracing genetic history of modern humans using X-chromosome lineages
Human Genetics (2007)
-
Genetic variability in a genomic region with long-range linkage disequilibrium reveals traces of a bottleneck in the history of the European population
Human Genetics (2005)